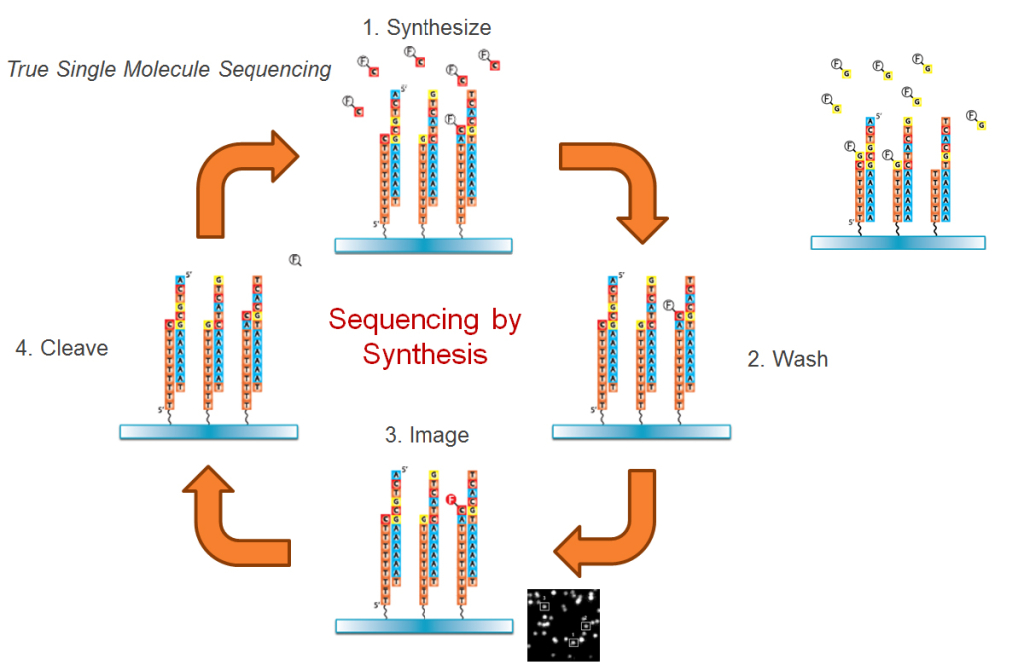
 DNA sequencing has undergone a revolution within the last fifty years. Beginning in the late 1970s, researchers have been improving the way we sequence DNA and trying to provide accurate results at a higher throughput. The first generation of DNA sequencing technology, Sanger Sequencing, involved adhering nucleotides (dNTPs) and modified nucleotides (ddNTPs) to a template strand of DNA that was later run through gel electrophoresis to determine the exact order of the sequence. While this method of sequencing is accurate, it can be slow and extremely costly, keeping many smaller research labs from being able to run tests that could further their studies. Though this technique is still a valid way to sequence, scientists quickly began looking for technology that would allow for achieving faster and cheaper results. To accomplish this, numerous companies developed second and third generation methods of sequencing. Among these companies producing third generation sequencing technologies was Helicos Biosciences with their Heliscope Genetic Analysis System; the first commercially available single molecule sequencer. At it’s core, the HeliScope relies on True Single Molecule Sequencing (tSMS™) to improve upon the methods developed for first-generation sequencing. The tSMS™ technology, employed at SeqLL, differs from previous methods in that it sequences strands in a massively parallel manner without any amplification steps, meaning that many strands can be sequenced at once and also provide unique information. The HeliScopes are able to read flow cells that are packed very densely with genetic material, allowing for higher throughput. The tSMS™ method typically utilizes a dT50 primer to initiate sequencing from a 3′ poly-A tail, though other capture primers may be used to increase the specificity of sample hybridization. When the strand is adhered to the flow cell, a laser illuminates the field, causing the incorporated “Virtual Terminators” to fluoresce. A CCD camera then takes pictures of the flow cell, marking the location of each strand. When the incorporation chemistry is complete, scientists and informatics experts work to align the sequence information prior to distributing these alignments to researchers and customers. The tSMS™ method is advantageous as it allows researchers to directly sequence the DNA without sample preparation steps like ligation or PCR amplification. Furthermore, the technology allows for massively parallel sequencing, increasing the throughput tremendously without requiring phased incorporations. References:
DNA sequencing has undergone a revolution within the last fifty years. Beginning in the late 1970s, researchers have been improving the way we sequence DNA and trying to provide accurate results at a higher throughput. The first generation of DNA sequencing technology, Sanger Sequencing, involved adhering nucleotides (dNTPs) and modified nucleotides (ddNTPs) to a template strand of DNA that was later run through gel electrophoresis to determine the exact order of the sequence. While this method of sequencing is accurate, it can be slow and extremely costly, keeping many smaller research labs from being able to run tests that could further their studies. Though this technique is still a valid way to sequence, scientists quickly began looking for technology that would allow for achieving faster and cheaper results. To accomplish this, numerous companies developed second and third generation methods of sequencing. Among these companies producing third generation sequencing technologies was Helicos Biosciences with their Heliscope Genetic Analysis System; the first commercially available single molecule sequencer. At it’s core, the HeliScope relies on True Single Molecule Sequencing (tSMS™) to improve upon the methods developed for first-generation sequencing. The tSMS™ technology, employed at SeqLL, differs from previous methods in that it sequences strands in a massively parallel manner without any amplification steps, meaning that many strands can be sequenced at once and also provide unique information. The HeliScopes are able to read flow cells that are packed very densely with genetic material, allowing for higher throughput. The tSMS™ method typically utilizes a dT50 primer to initiate sequencing from a 3′ poly-A tail, though other capture primers may be used to increase the specificity of sample hybridization. When the strand is adhered to the flow cell, a laser illuminates the field, causing the incorporated “Virtual Terminators” to fluoresce. A CCD camera then takes pictures of the flow cell, marking the location of each strand. When the incorporation chemistry is complete, scientists and informatics experts work to align the sequence information prior to distributing these alignments to researchers and customers. The tSMS™ method is advantageous as it allows researchers to directly sequence the DNA without sample preparation steps like ligation or PCR amplification. Furthermore, the technology allows for massively parallel sequencing, increasing the throughput tremendously without requiring phased incorporations. References:
- http://krishna.gs.washington.edu/GS_373_12/lec_3_1.pdf
- http://www.ncbi.nlm.nih.gov/pubmed/20890904
- http://www.nature.com.ezproxy.bu.edu/nbt/journal/v27/n9/full/nbt.1561.html
- http://www.fiercebiotech.com/story/troubled-sequencing-pioneer-helicos-shelters-bankruptcy-court/2012-11-19
- http://www.researchgate.net/publication/44802897_Single-molecule_sequencing_sequence_methods_to_enable_accurate_quantitation/file/5046351794b1072287.pdf
- http://genome.wustl.edu/articles/detail/dna-sequencing-technology-a-perspective-from-dr-elaine-mardis/
- http://hmg.oxfordjournals.org/content/19/R2/R227.long


Leave A Comment